October is Breast Cancer Awareness Month, a time to honor the patients and their families worldwide whose lives have been impacted by breast cancer and to recognize the progress being made to combat this pervasive disease.
Globally, breast cancer remains the most common cancer among women, with over 2.3 million new cases reported in 2022.1 In 2025, in the United States alone, an estimated 316,950 women and 2,800 men will be diagnosed2 with invasive breast cancer. Among these patients, about 70%3 – more than 225,000 – will be diagnosed with hormone receptor-positive (HR+) breast cancer. Given these alarming statistics, the need for innovative approaches to fighting breast cancer has never been more pressing.
Genomic Testing
Genomic Testing: Unlocking the Power of Truly Personalized Breast Cancer Treatment
Historically, endocrine therapy and chemotherapy have been the standard of care for HR+breast cancer. However, one of the most transformative advances of the past two decades has been the rise of genomic testing – including the Prosigna® Breast Cancer Assay. These tests can identify patients who may benefit from more intensive therapy or safely avoid chemotherapy – and its long-term toxicities – while still helping patients remain cancer-free. Data strongly support their use in postmenopausal, low-risk, node-negative patients, but evidence has been less definitive for premenopausal women and high-risk postmenopausal patients, particularly when predicting chemotherapy benefit.
To address this challenge, the National Institute for Health and Care Research and the University College London in the United Kingdom (UK) have sought to answer the question of whether these high-risk patients can safely avoid chemotherapy through a series of prospective trials called OPTIMA:4 Optimal Personalised Treatment of Early Breast Cancer Using Multi-parameter Analysis.
Evidence in Action
What We’re Learning from the OPTIMA Trials
OPTIMA Prelim Trial: Prosigna Selected for Best Value of Information
In 2014, a preliminary trial tested the feasibility of enrolling patients in a randomized prospective study to evaluate whether a biomarker could guide more accurate use of chemotherapy in high-risk patients. Multiple risk stratification tests were run on these patient samples, and the results were subjected to a “Value of Information” analysis to determine which test to evaluate in a larger, prospective trial. The Prosigna test was ultimately selected as the enrollment test for the OPTIMA main trial, a 4,500 patient, prospective study.
In May 2025, the long-term outcomes of this study were presented5 at the ESMO Breast Cancer Congress,6 with the following key findings:
- The Prosigna test demonstrated higher prognostic accuracy in high-risk patients (node-positive and high-risk node-negative) compared to the test originally used to assign treatment groups.
- Among the patients with low recurrence scores (≤ 25), 22% had discordantly high Prosigna scores (≥ 60). This group had the worst 10-year recurrence free interval (RFI) at 74% (95% Confidence Interval [CI] 62-83%), significantly worse than the concordantly low-risk group (RFI 89%, 95% CI 84-92%; Hazard Ratio [HR] 2.32, 95% CI 1.26-4.24, p = 0.0067; adjusted HR 1.97, 95% CI 1.01-1.97).
- These results confirm the superior prognostic performance of the Prosigna test, in line with previous retrospective studies.
OPTIMA Main Trial: Moving from Assumptions to Answers
OPTIMA main is a prospective, randomized, blinded, Phase 3 trial. The trial is evaluating whether outcomes are similar when patients identified as having low-risk disease by a genomic test – the Prosigna test – are treated with hormonal therapy alone versus the combination of hormonal therapy with chemotherapy.
Since 2017, the OPTIMA main trial has enrolled more than 4,400 patients across the UK, Norway, Australia, New Zealand, and Thailand to compare standard of care endocrine therapy and chemotherapy to therapy guided by the Prosigna test. Enrollment was completed in the UK in late 2024 and in Norway in August 2025 and is set to wrap up in the remaining countries by the end of 2025. The primary analysis of the results, expected in 2026, aim to deliver the definitive answer on whether the Prosigna test can predict which clinically high-risk patients benefit from chemotherapy, and which can safely forego it.
OPTIMA-Young Trial: Empowering Confident Decisions for Premenopausal Women
OPTIMA-Young is a global trial focused entirely on premenopausal women, spanning 15 countries across Europe, Oceania, and Latin America with a target enrollment of 3,380 patients. Running in tandem with the OPTIMA main trial, OPTIMA-Young will contribute to a combined analysis encompassing data from 5,000 premenopausal participants across both studies. Mirroring the OPTIMA main trial design, it uses the Prosigna Breast Cancer Assay to guide treatment decisions and will strengthen the data in premenopausal women.
The Prosigna Test: The Test To Answer More Than One Question
The Prosigna Breast Cancer Assay is an advanced genomic test for patients with invasive, early-stage HR+ breast cancer that helps guide more personalized treatment decisions. It is the only assay validated using the four intrinsic molecular subtypes – Luminal A, Luminal B, HER2-enriched, and Basal-like.
The Prosigna test uniquely integrates this subtype information with a proliferation score and key clinicopathologic factors to generate a single 10-year Risk of Recurrence (ROR) score. This insight into the likelihood of cancer returning empowers physicians and patients to make more precise, personalized treatment decisions to help lead to better outcomes.
Looking Ahead
Advancing the Future of Breast Cancer Care Together
Breast Cancer Awareness Month offers us all a moment to take stock of how far we’ve come, and how much further we have yet to go. Standard treatments like endocrine therapy and chemotherapy have saved countless lives, but advances in genomic testing, like the Prosigna Breast Cancer Assay, and discoveries made in large-scale studies such as the OPTIMA trials, are opening the door to truly personalized care in breast cancer.
By combining awareness with continued research and innovation, we move closer to a future where physicians have the data they need to confidently guide treatment decisions – ensuring every patient receives the right therapy at the right time, with the best chance for long-term health. Veracyte stands alongside physicians, patients, and the broader community as an ally to not only raise awareness, but to also recognize the progress transforming breast cancer care for patients everywhere.
Talk to your doctor about whether the Prosigna test is right for you.
If you have questions or need more information about the Prosigna test, we’re here to help. Connect with a member of our team at [email protected].
September is Prostate Cancer Awareness Month, a dedicated time to recognize the brave individuals impacted by the disease and honor the ongoing research and innovation reshaping patient outcomes.
Nearly 314,000 patients in the United States will be diagnosed with prostate cancer this year, and the disease will claim approximately 35,770 lives, according to the American Cancer Society. In addition, the rate of men diagnosed1 with advanced disease has been growing in recent years.
What is Prostate Cancer?
Prostate cancer occurs when cells in the prostate gland start to grow abnormally. These abnormal cells can form tumors and spread to other parts of the body. Even though certain prostate cancers can grow rapidly, the majority advance slowly and respond well to treatment .
While anyone with a prostate may develop prostate cancer, some people face higher risk due to specific factors.2 African American men in the United States are 1.7 times more likely3 to develop prostate cancer compared to European American men. The disease occurs less often2 in Asian American, Hispanic, and Latino men. While rare in men younger than 40, the chance of having prostate cancer rises rapidly after 50, with about 6 in 10 prostate cancers2 found in men older than 65. Although the majority of cases have no family history, a man’s risk more than doubles2 if a close relative has had the disease.
Screening and Diagnosing Prostate Cancer
While symptoms of early prostate cancer are rare, they can include problems urinating and blood in the urine or semen. As the cancer advances and spreads, symptoms can include erectile dysfunction, pain in different parts of the body, weakness or numbness in the legs or feet, loss of bladder control, and fatigue.
To screen for prostate cancer, clinicians typically use a PSA (prostate-specific antigen) blood test, with or without a digital rectal exam (DRE). If cancer is suspected, additional testing is performed such as an MRI and prostate biopsy. The biopsy tissue is then graded using the Gleason score (Grade Group). Clinical stage is determined by combining PSA, biopsy results, and imaging findings. These imaging scans may also be used to evaluate whether the cancer has spread beyond the prostate (metastasis).
Learn more about when screening for prostate cancer is recommended.
The Challenge in Prostate Cancer Management
Prostate cancer is heterogeneous, meaning it doesn’t behave the same way in every person. Some prostate cancers grow very slowly, while others are aggressive and can spread quickly, making clinical decision-making challenging.
Historically, decisions were primarily made based on clinical and pathological factors, however, these factors alone do not always reflect the true biology of the tumor, which can lead to over- and under-treatment.
The good news is that gene expression testing is enabling clinicians and patients to better understand the unique biological characteristics of the disease and make more informed treatment decisions.
Genomic Technology as a Better Guide
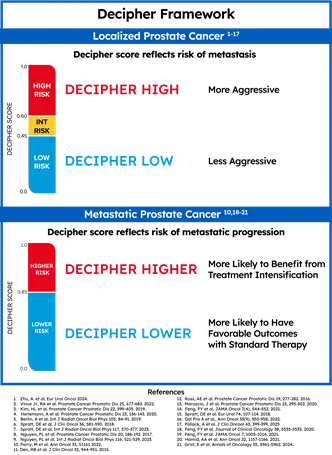
Looking at the activity of certain genes in tumor cells using molecular tests like the Decipher Prostate Genomic Classifier can often better predict how likely a cancer will grow or spread.
Decipher Prostate delivers trusted genomic insights to help clinicians and patients make confident, personalized treatment decisions across the prostate cancer continuum. The test is a highly accurate predictor of metastasis, prostate cancer-specific mortality, and overall survival in localized patients, among other endpoints. Additionally, for patients with metastatic prostate cancer, the Decipher score indicates the likelihood of cancer progression and survival benefit with treatment intensification.
What is the Decipher Prostate Test?
Decipher Prostate is a 22-gene genomic test that looks at the molecular characteristics of the prostate cancer tumor to assess its aggressiveness, or likelihood of spreading outside the prostate, providing actionable information for treatment decision-making across the spectrum of prostate cancer – post-biopsy, post-RP (radical prostatectomy), and advanced disease. It is performed on tissue collected during a patient’s existing prostate tumor biopsy or radical prostatectomy.
The result is a number from 0-1 (the patient’s Decipher score) classified as Decipher Low Risk, Decipher Intermediate Risk, or Decipher High Risk. For patients with localized or regional prostate cancer, a Decipher score indicates a patient’s risk of metastasis, helping to determine treatment timing and intensity. For patients with metastatic prostate cancer, the Decipher score indicates the likelihood of cancer progression and survival benefit with treatment intensification.
Patient Journeys: A Urologist’s Experience
Urologist Dr. Arnold Bullock, from the Washington University School of Medicine, shares his very personal story of being diagnosed with prostate cancer and using the Decipher Prostate test in his clinical practice for his patients. Watch his conversation with urologist and former colleague, Dr. Eric Kim, a key opinion leader from the University of Nevada, Reno School of Medicine.
Building Clinical Value and Rigorous Evidence
Decipher Prostate is the most extensively validated gene expression test in prostate cancer, with evidence from over 200,000 patients, more than 25 prospective randomized clinical trials, and upwards of 95 peer-reviewed publications demonstrating its clinical validity and utility.
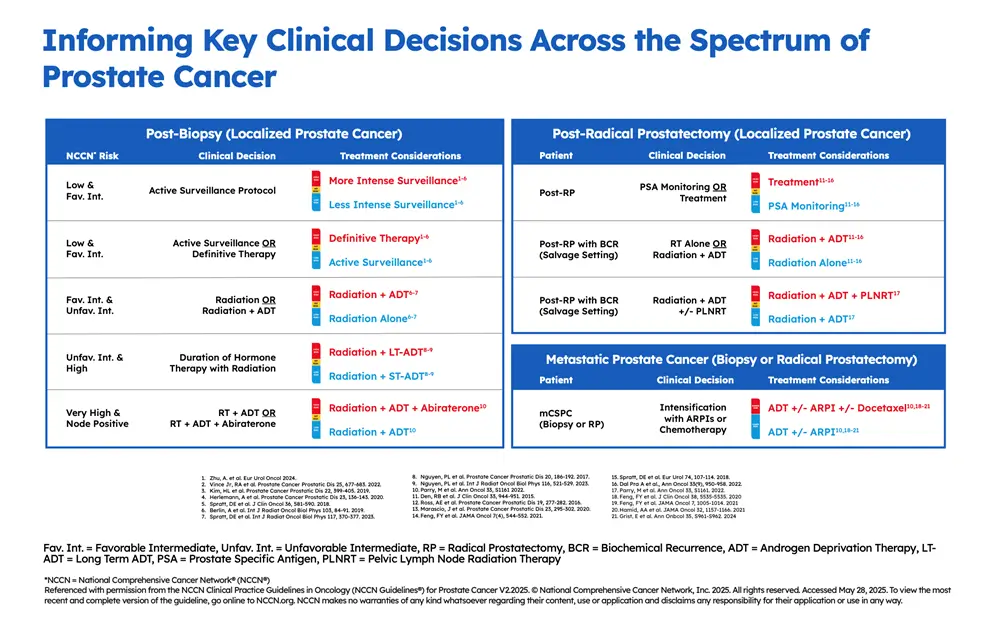
It is the only gene expression test to achieve Simon Level I evidence status and inclusion in the risk-stratification table in the most recent version of the NCCN Clinical Practice Guidelines In Oncology (NCCN Guidelines®)4 for Prostate Cancer.
Recent Publication Highlight
Findings suggest Decipher Prostate can help clinicians better personalize chemotherapy decisions for patients with metastatic prostate cancer New data published online in Cell, from the randomized, prospective, Phase 3 STAMPEDE trial,5 show that Decipher Prostate predicts which patients with metastatic cancer are likely to benefit from treatment intensification with the chemotherapy docetaxel and which are not likely to benefit and can therefore avoid its unnecessary toxicity. Read Press Release.
What’s Next: Personalized Prostate Cancer Treatment Planning
As of June 2025, Decipher Prostate expanded into metastatic prostate cancer with the Decipher Prostate Metastatic Genomic Classifier. Approximately 10% (or about 30,000)1 of all prostate cancers diagnosed annually in the United States are metastatic, and Veracyte believes this important new tool will help clinicians make better-informed treatment recommendations.
Today, Veracyte estimates1 that about 40% of patients in the US with prostate cancer receive molecular testing to help guide their care, and as more patients embrace earlier testing, the opportunity for timely diagnosis can offer the potential to halt disease progression before it reaches advanced stages and improve long-term quality of life.
At the end of this month, multiple abstracts will be presented at ASTRO 2025 demonstrating Veracyte’s commitment to continue working with research collaborators to advance our collective knowledge of prostate cancer.
Throughout the prostate cancer care continuum, both clinicians and patients must navigate complex treatment planning decisions. Advanced genomic tests, like Decipher Prostate, capable of providing whole-transcriptome data and deeper molecular insights, will enable more precise, effective treatments for better patient outcomes.
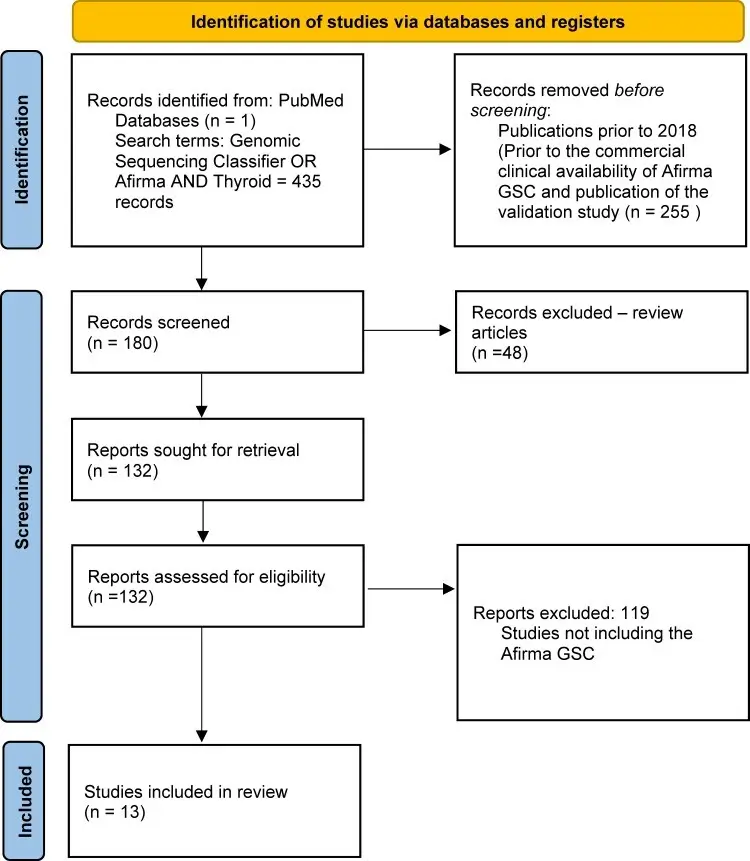
Afirma GSC Shows Value in Ruling Out Malignancy in Indeterminate Thyroid Nodules (ITN)
An indeterminate thyroid nodule (ITN) carries a ~25% risk of malignancy (ROM), and historically diagnostic surgery for ITNs revealed a benign histopathology in 75% of cases.1
Molecular testing attempts to help prevent unnecessary surgeries with a focus on “rule-out” metrics such as Sensitivity (SN) and Negative Predictive Value (NPV). Tests may be optimized to keep the false negative rate low in validation studies, but how does that translate into real-world outcomes?
Ahmadi et al. sought to evaluate the performance of Afirma GSC’s rule-out capabilities in their evaluation of over 800 Bethesda III/IV nodules, with a median follow-up time of 23 months.
In this independent, multicenter study, the authors sought to understand key performance metrics, including SN and NPV in their evaluation of real-world patient outcomes.
Afirma GSC Demonstrated Strong Performance in Line with Validation Study Metrics
When stratifying performance metrics between Bethesda III and Bethesda IV nodules, Afirma GSC performed similarly across these two nodule types. While some studies may have a limited number of higher-risk Bethesda IV nodules included in the cohort,2 a total of 215 Bethesda IV nodules were evaluated by Ahmadi et al. Notably, the SN and NPV performance were in line with or above the validation study.3
Given that 80% of patients with Bethesda IV cytology who do not undergo molecular testing end up having surgery,4 Afirma GSC can confidently identify benign nodules in Bethesda IV cytology to help prevent unnecessary surgeries for these patients.
Ahmadi et al. Found That Afirma Demonstrated a False Negative Rate of 1.5%
When looking at performance among Afirma GSC Benign nodules, 528 patients received an Afirma GSC Benign result for a Benign Call Rate (BCR) of 63% overall. With the majority undergoing surveillance (N = 443) and some patients opting for surgery – including delayed surgery – the overall false negative rate is 1.5% (8/528 nodules) when assuming the unoperated Afirma GSC Benign nodules would have been benign had they been removed.
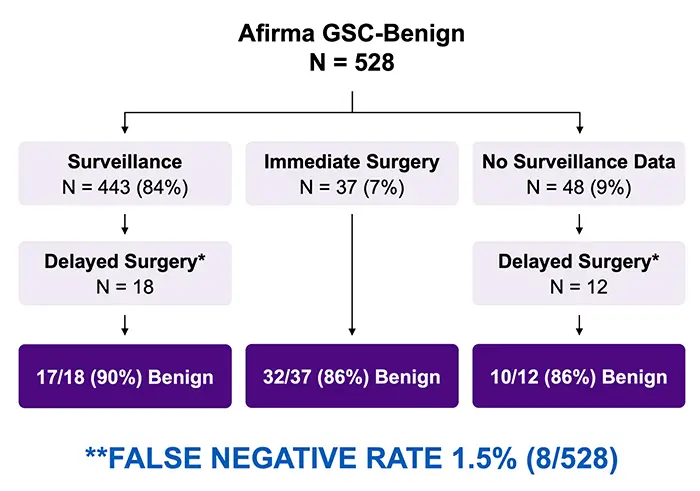

Clinical Management of Afirma GSC Benign Nodules
Ultimately, Ahmadi et al. found that Afirma GSC can be a key tool for managing patients with indeterminate cytology in a variety of practice settings and prevalence of malignancy. The majority of GSC-benign nodules remained stable over the surveillance period, and the authors found that “GSC-benign thyroid nodules can be observed similarly to thyroid nodules with benign cytology”,5 reinforcing Afirma’s “rule-out” capabilities.
May is Bladder Cancer Awareness Month—a meaningful time to spotlight a disease that will be diagnosed in nearly 85,000 people in the U.S. this year alone and is expected to claim more than 17,000 lives, according to the American Cancer Society.1
Unfortunately, the bladder cancer patient journey is rarely straightforward. From vague initial symptoms to high rates of disease recurrence and challenging treatment decisions, patients and clinicians alike often face uncertainty at nearly every stage.
Fortunately, advances in genomic technology now offer new tools to help clinicians and their patients make better-informed and more-confident treatment decisions based on the biology of each patient’s cancer.
Understanding Bladder Cancer
Bladder cancer most commonly affects older adults, with 90% of cases2 occurring in people over age 55 and the average age at diagnosis being 73. Smoking is the most common risk factor. While bladder cancer’s symptoms—such as blood in the urine, increased urination frequency or urgency, or pain during urination—can be mistaken for other conditions, early diagnosis is important.
Diagnosis of bladder cancer typically involves cystoscopy, in which a camera is used to visualize the inside of the bladder. If cancer is suspected, generally patients are offered a transurethral resection of a bladder tumor (TURBT), a procedure that is both diagnostic and therapeutic. Once definitively diagnosed, patients are categorized as having:
- Non-muscle-invasive bladder cancer (NMIBC) – where cancer is confined to the inner lining of the bladder (about 70% of cases3); or
- Muscle-invasive bladder cancer (MIBC) – where cancer has penetrated the muscle wall or beyond (about 30% of cases).
MIBC is considered more aggressive and requires urgent intervention. However, both NMIBC and MIBC have high rates of recurrence, with the cancer typically requiring lifelong follow-up care.
Navigating a Complex Treatment Journey
Treatment of NMIBC and MIBC differ significantly and each typically involves multiple steps.
- NMIBC treatment often includes intravesical therapy, in which a liquid drug – typically the immunotherapy Bacillus Calmette-Guerin (BCG) or a chemotherapy cocktail (gemcitabine and docetaxel) – is delivered directly into the bladder. If patients respond well to intravesical therapy, they will typically receive maintenance treatment for 1-3 years. If they do not respond well to intravesical therapy, clinical guidelines recommend radical cystectomy (RC) or chemotherapy plus radiation; other options include RC, systemic chemotherapy, immunotherapy, or intravesical investigational drugs.
- MIBC treatment typically involves neoadjuvant chemotherapy (NAC) followed by radical cystectomy or a combination of systemic chemotherapy and radiation. In some cases, additional chemotherapy or immunotherapy may be administered after surgery or radiation.
Treatment decisions to date have been primarily guided by the cancer’s grade and stage, which are informed by histopathology and imaging scans. These clinical risk factors, however, can be imprecise, which can lead to patients being overtreated or undertreated. In fact, data show that approximately 40% of patients with NMIBC are upstaged at radical cystectomy4 to MIBC. The growing number of treatment options available further adds to the complexity.
Genomic Technology as a Better Guide
This is where advanced genomic technology comes in.
The Decipher Bladder Genomic Subtyping Classifier is a clinically available genomic test that provides a deep biological characterization of each patient’s bladder tumor to help guide treatment decisions for patients with NMIBC or MIBC. Developed through RNA whole-transcriptome analysis and machine learning, the 219-gene test classifies bladder tumors into five distinct molecular subtypes—each with unique biology and potential treatment implications. This information can help guide decisions at key moments in a patient’s care journey.
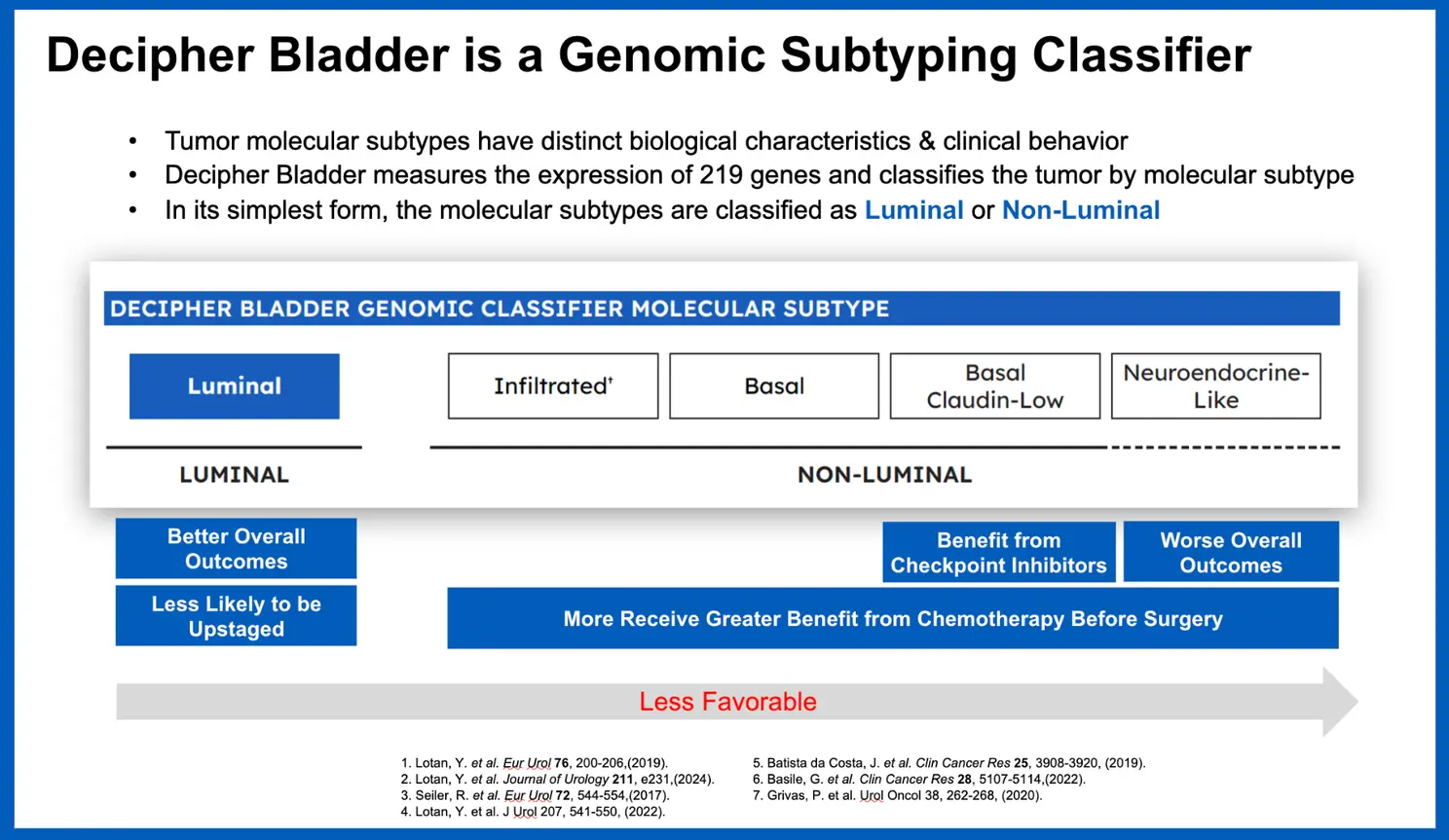

Building Rigorous Clinical Evidence
The Decipher Bladder test is supported by a dozen published studies demonstrating its performance and clinical utility. Such clinical evidence has led to the test’s coverage by Medicare and growing interest among clinicians.
Just recently, a study published in European Urology Open Science showed that the test can identify patients whose disease may be more aggressive than initially staged via TURBT.
Additionally, through its whole-transcriptome-based Decipher GRID (Genomic Resource for Intelligent Discovery) research tool, Veracyte and its collaborators are deriving additional insights that may ultimately impact patient care. For example, among nearly 20 abstracts presented on our Decipher Prostate and Bladder tests at the recent American Urological Association annual meeting, researchers used Decipher Bladder GRID to identify a subset of patients with high-risk NMIBC who responded better to therapy with sequential intravesical gemcitabine and docetaxel (Gem/Doce), compared to guideline-recommended intravesical Bacillus Calmette-Guerin (BCG), based on their tumors’ underlying biology. Such findings could be particularly helpful, given ongoing supply challenges with BCG.
What’s Next: Bladder Cancer MRD Testing
After completing curative treatment—such as surgery, chemotherapy, or radiation—clinicians and patients are often left with lingering questions such as: Has the cancer truly been eliminated? Is more treatment needed? Traditional imaging and pathology often can’t detect small amounts of residual disease, which can lead to over-, under- or delayed treatment —all of which can negatively impact patient outcomes.
To address this, Veracyte is developing a novel minimal residual disease (MRD) test for MIBC, which is performed on a patient’s blood sample following treatment. The test uses a combination of whole-genome sequencing and artificial intelligence (AI) to enable earlier detection and improved outcomes throughout the patient’s treatment and follow-up care.
In a podium presentation at the recent European Association of Urology Congress (EAU25), investigators shared promising findings from the large, independent, interventional TOMBOLA trial, which showed that Veracyte’s MRD testing platform detected cancer in patients treated for MIBC with more accuracy than digital droplet PCR and approximately three months earlier than standard imaging techniques.
At every stage of the bladder cancer journey, clinicians and patients face difficult decisions—often with limited information. Genomic advances like the Decipher Bladder test are helping to change that by delivering meaningful biological insights that can lead to more personalized, informed, and effective care. Because better insights lead to better outcomes.
An indeterminate thyroid nodule (ITN) carries a ~25% risk of malignancy (ROM),1 and molecular diagnostics can play a key role in ruling out cancer for the majority of ITN nodules that are benign. Molecular diagnostic tests are designed to confidently rule out malignancy in these nodules using analytical and clinical validation studies, but not all tests perform the same in a real-world setting.
Published studies of single institution experiences with a test may not be representative of the broader performance. Aggregating single-center experiences into a meta-analysis can robustly compare clinical validation (CV) metrics versus real-world performance.
Nasr et al. performed a meta-analysis of Afirma GSC, evaluating nearly 2000 ITNs across 13 sites to compare the clinical performance of Sensitivity (SN), Specificity (SP), Positive Predictive Value (PPV), and Negative Predictive Value (NPV) to the validation study.2 While other tests have multiple institutional experience studies, Nasr et al. performed the meta-analysis aligning rules and assumptions to evaluate the data in a uniform method.
When compared to the clinical validation study, the meta-analysis found the real-world experience matched or exceeded the metrics from the CV. Notably, statistically significant improvements were seen in Specificity, PPV, NPV, and the benign call rate (BCR):
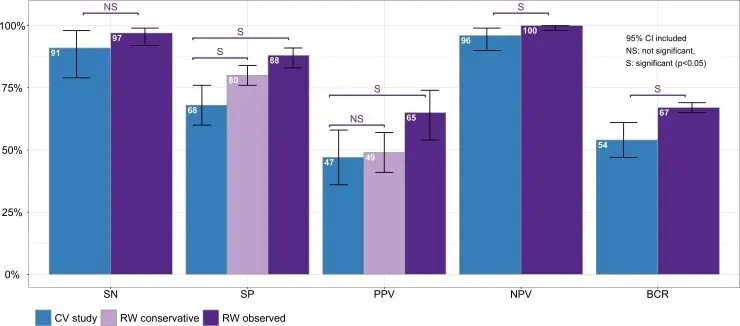
As most patients with an ITN are looking for molecular testing to guide the decision to choose surveillance over surgery, the improved Sensitivity and significant improvement in BCR demonstrate the value of an Afirma Benign result. Across all sites included in the final data set, the BCR and Sensitivity performed above or consistent with the validation study metrics. Consistent performance and high confidence in a rule-out result can be a critical component of a treatment decision, and the meta-analysis confirms the accuracy of an Afirma Benign result.
Afirma GSC utilizes a whole transcriptome-derived platform combined with machine learning. Using unique classifiers and a robust sample set for the clinical validation study, Afirma GSC’s real-world performance indicates the test continues to evolve and improve across a variety of clinical settings.
Previously we’ve discussed how cancer isn’t a single mutation event. Similarly, cancer behavior isn’t limited to cancer cell activity alone. This view is devoid of the context that surrounds and influences the cancer cells – known as the “tumor microenvironment,” or “TME.”
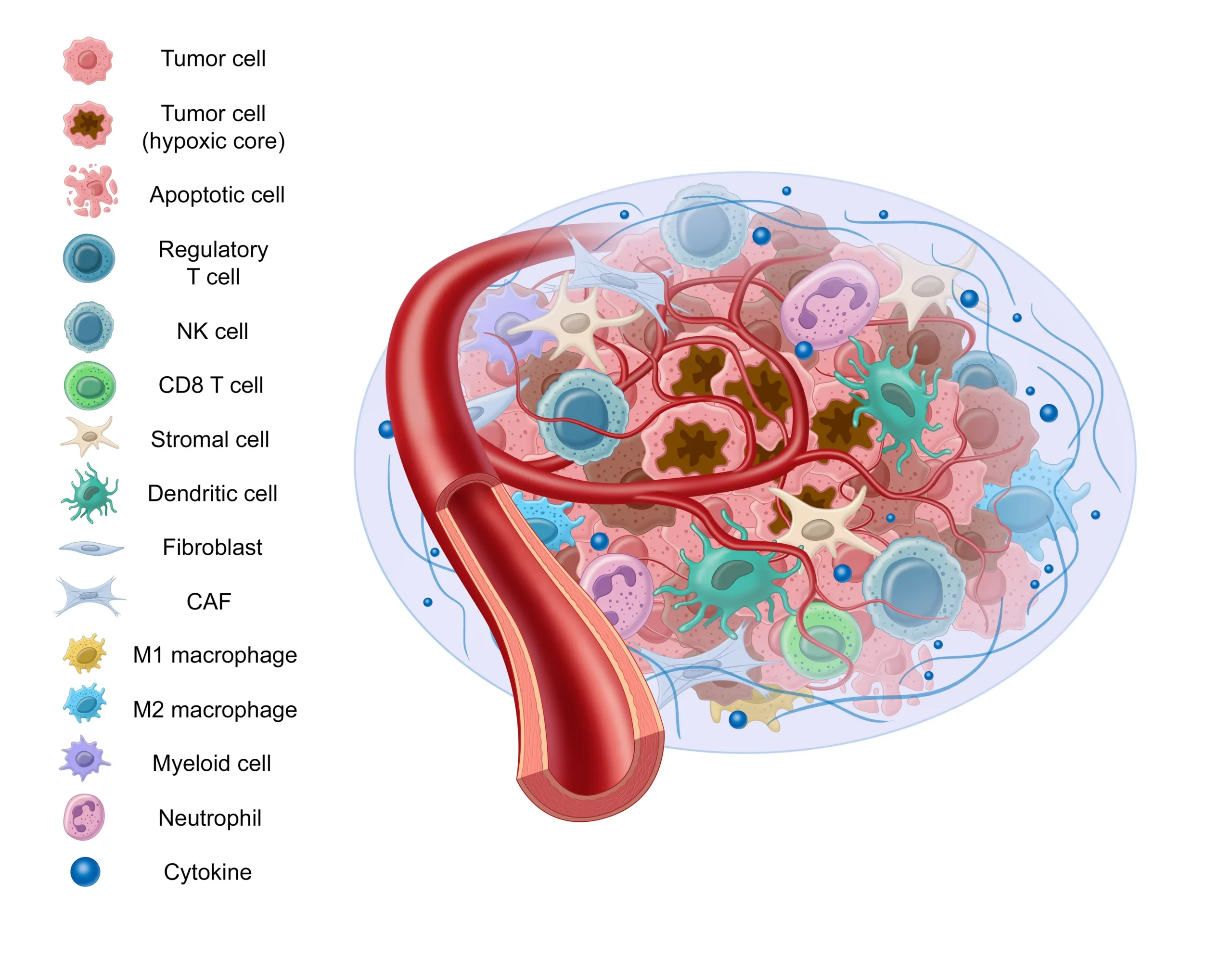
English surgeon and pathologist Stephen Paget proposed a “seed and soil” hypothesis which describes cancer cells as a seed and the tumor microenvironment as the soil. This hypothesis is now widely accepted and provides an apt analogy to help understand the role the tumor microenvironment plays in cancer progression.
There are many components to the tumor microenvironment – let’s review some of them:
Stromal Tissue
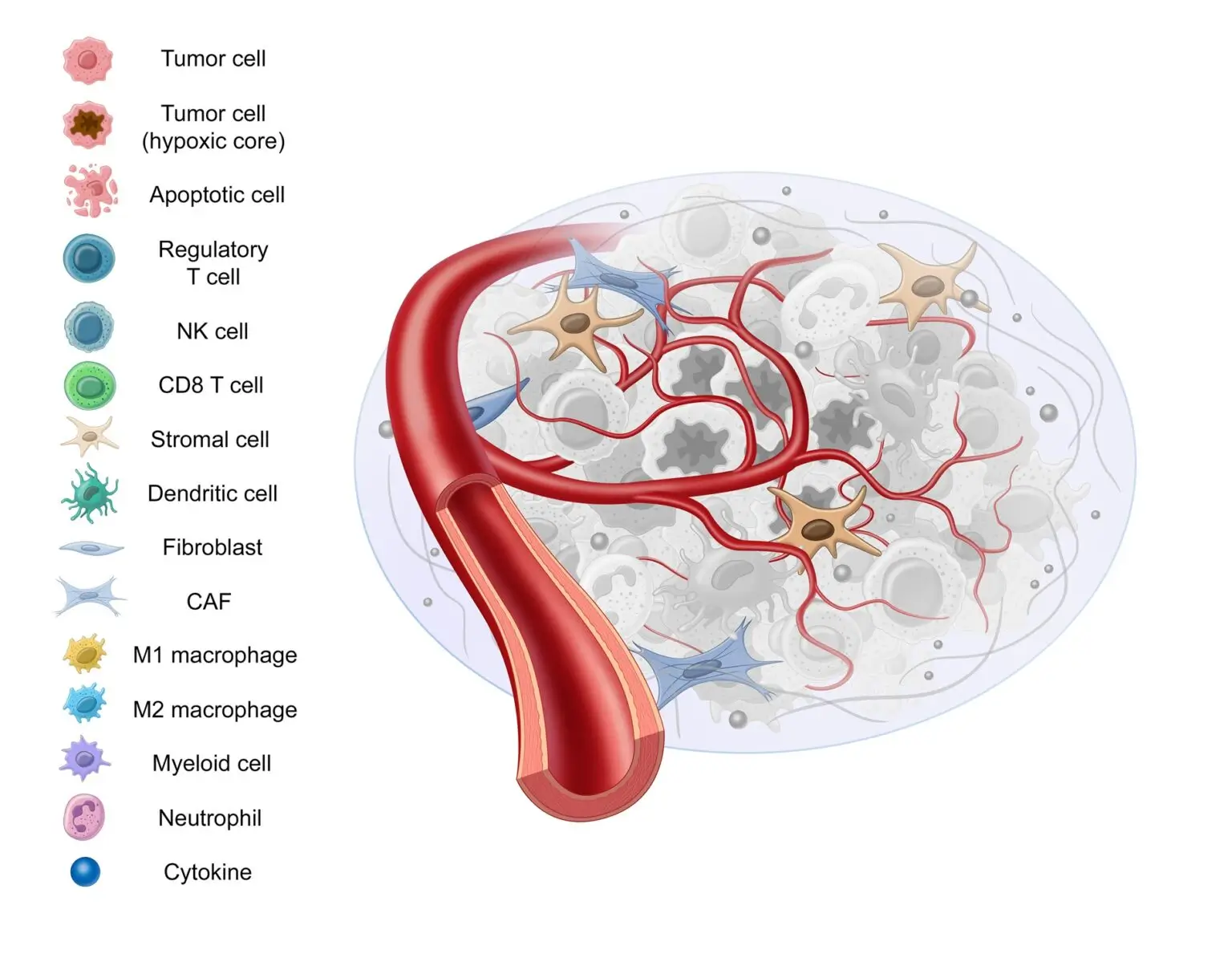
Stromal tissue is comprised of supportive and connective tissue, as well as fibroblasts, myofibroblasts, vascular cells, and macrophages, among others.
Stromal tissue doesn’t act as a bystander in the development and progression of cancer. It contributes to tumor growth by secreting tumor promoting factors, promoting angiogenesis, and altering fibroblasts, all of which we’ll cover in this blog post.1
Cancer Associated Fibroblasts
In a healthy environment, these fibroblasts play an important role in healing wounds – but in the presence of transforming growth factor-beta (TGF-β), they’re converted into cancer associated fibroblasts, or “CAFs.”2
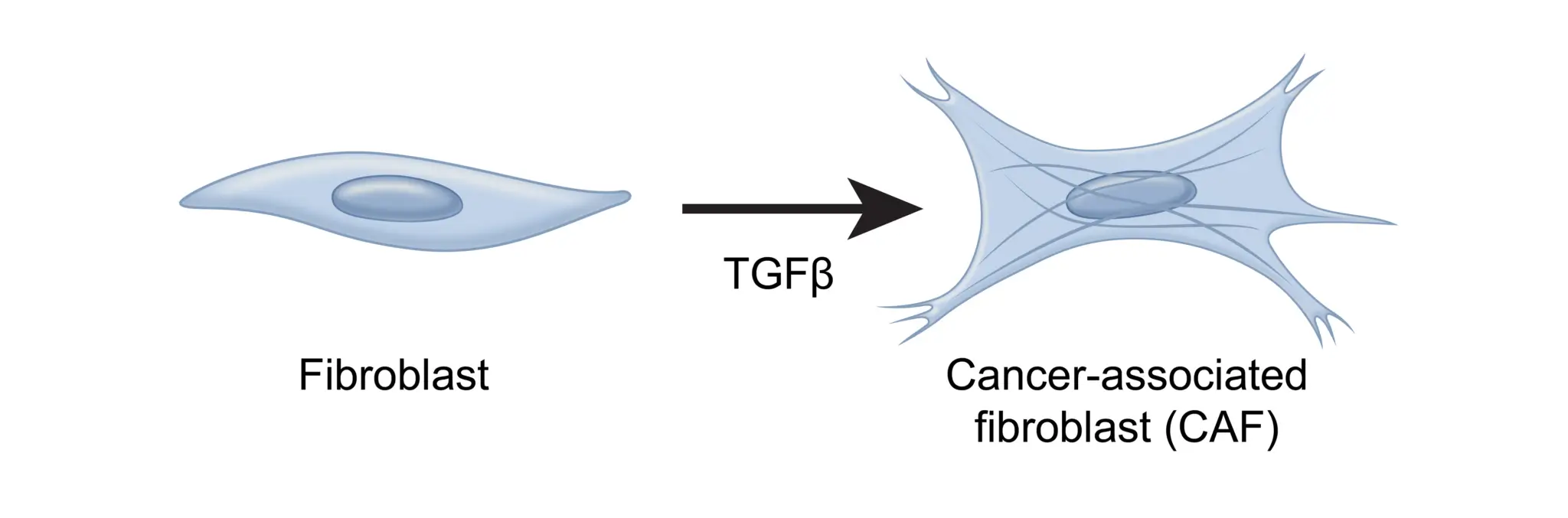
These CAFs are the most abundant cell type in the TME and have been identified as a predictor of aggressive thyroid cancer.3,4,5 Earlier this year at ENDO, research was presented regarding a novel thyroid-cancer specific CAF gene signature created using the Afirma exome-enriched molecular testing platform.5
CAFs promote tumor growth in a variety of ways, including remodeling the extracellular matrix to be more favorable to tumor growth or secreting cytokines that promote angiogenesis.3,6,7,8
Angiogenesis
Why is angiogenesis important? Blood vessels are needed to bring oxygen and nutrients to healthy, noncancerous cells, and cancer cells are no different.9 In order to grow and metastasize, new blood and lymphatic vessels must form from existing vessels. This process is known as “angiogenesis”.
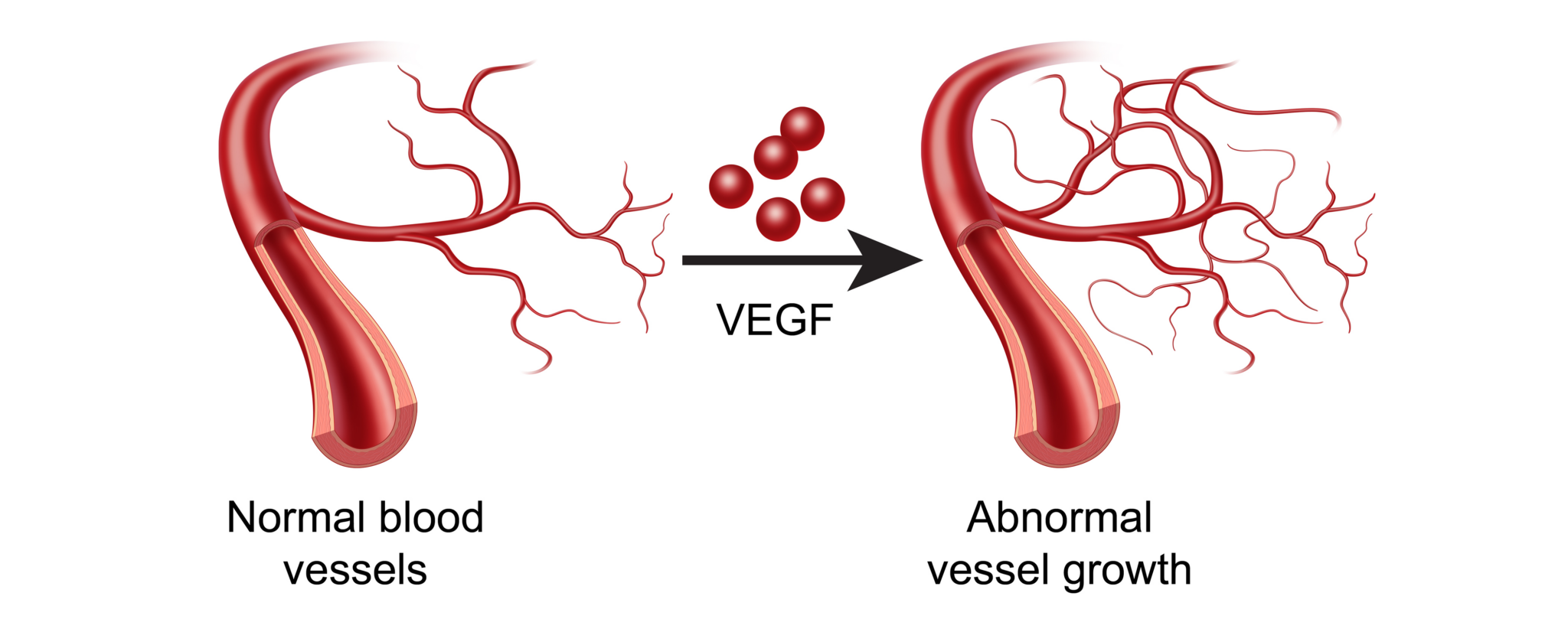
The difference in the TME is that these new vessels brought about by vascular endothelial growth factor (VEGF) or IL-8 are often too permeable, or “leaky”.10,11 Such vessels are detrimental to surrounding healthy tissue, but conducive to tumor growth.12
Hypoxia
Despite oxygen serving as a fundamental requirement for healthy cells, the tumor microenvironment is generally characterized by low levels of oxygenation, or “hypoxia.”9 Somewhere between 25 and 40% of invasive breast cancers, for instance, test positive for hypoxic markers.9
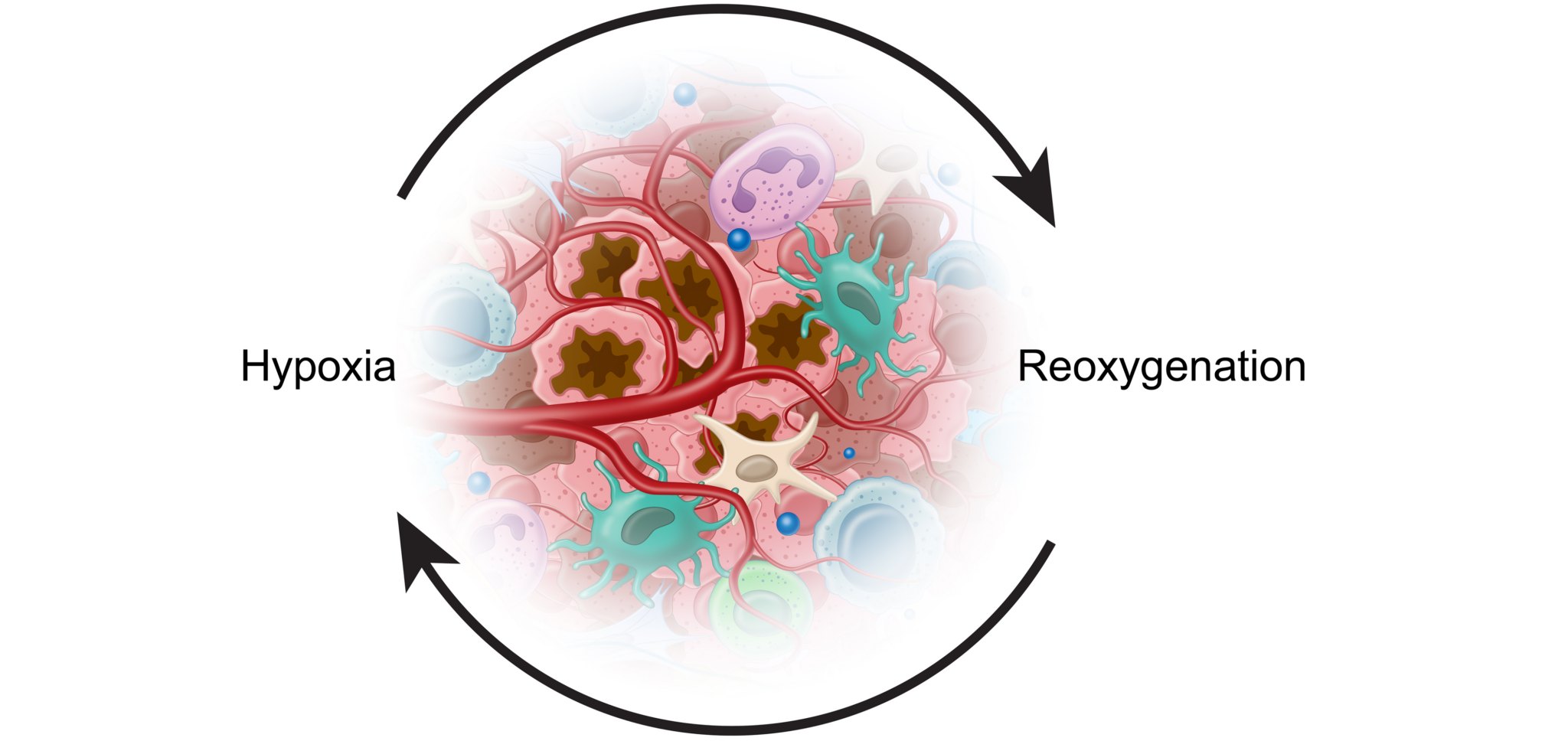
These periods of hypoxia are followed by periods of reoxygenation which lead to mutations, DNA strand breaks, mutation amplification, and other tumor-promoting complications.9,13 Cancer cells have adapted to these conditions and used them to their benefit.
The Warburg Effect
How are tumor cells able to survive and proliferate without oxygen? Healthy, normal cells use a process called Oxidative phosphorylation (OxPhos) to generate ATP, the energy for the cell. Without enough oxygen, this process can’t be completed. Cancer cells are able to switch from this process to glycolysis, which can be completed in the absence of oxygen, in order to produce energy.12,15 This is referred to as the “Warburg Effect”.
Interestingly, it is this same phenomenon that allows tumors to be detected by PET scans.16 Glycolysis uses significantly more glucose than non-cancerous cells, so a radioactive glucose analog is injected during a PET scan, which then accumulates in cancerous areas and results in visible “hot spots” in the scan.
Immune Cells
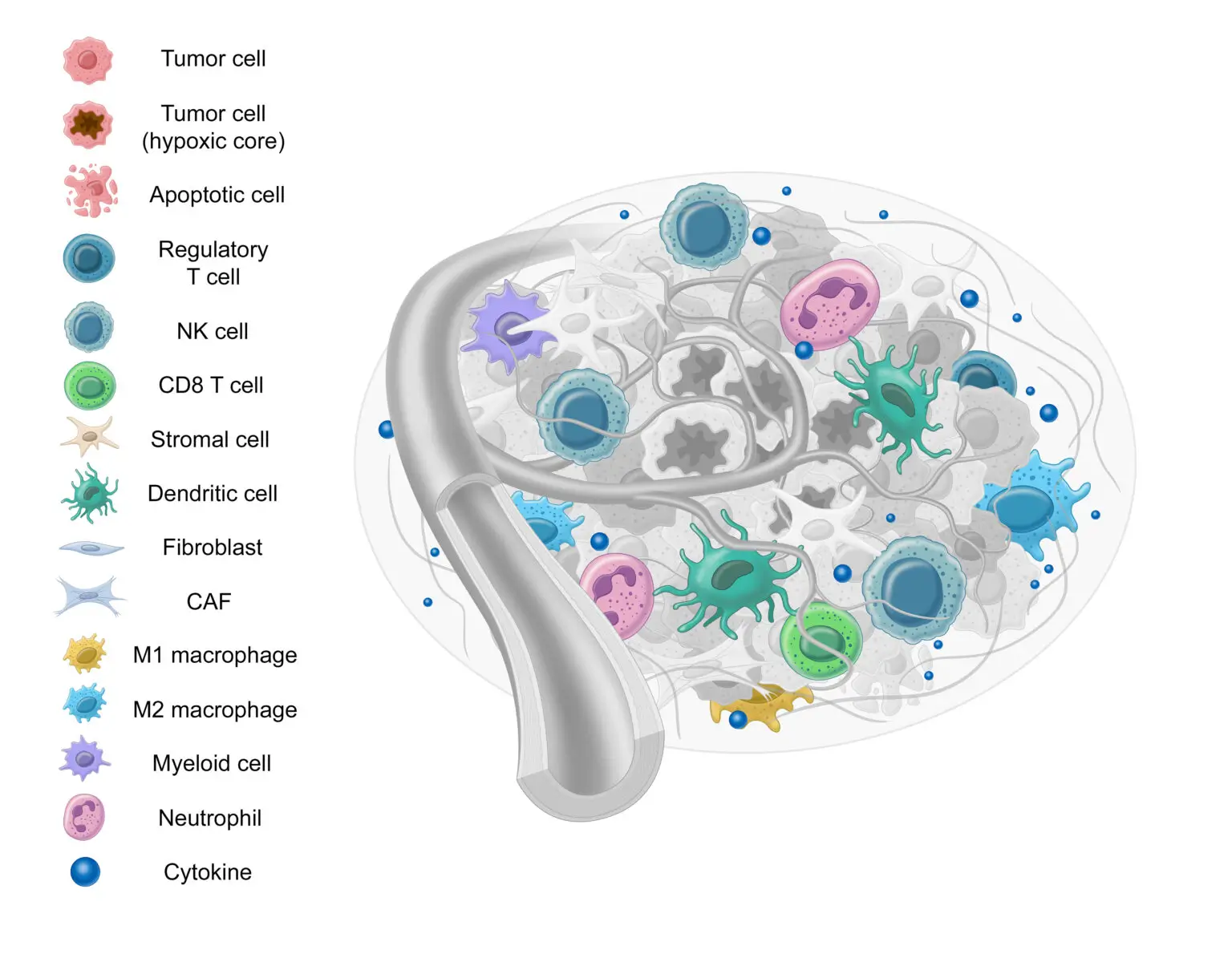
Immune cells serve a significant role in tumor progression and suppression.17-19 There are many different types of immune cells that interact with the TME, but here we’ll review the M1 and M2 macrophages, PDL1, CD4+ T cells, and CD8+ T cells.
Macrophages
First, let’s examine macrophages. Macrophages, specialized white blood cells that play a critical role in immune response, may also at times benefit the tumor microenvironment.10,21
The M2/M1 Macrophage Ratio
Due to their opposing nature, the ratio between M1 and M2 macrophages (or M1/M2 ratio) can provide potential insight into tumor progression.23,24 While unexplored in thyroid cancer, a lower M1/M2 ratio is correlated with poor outcomes in ovarian cancer, multiple myeloma, pediatric classical Hodgkin lymphoma, colorectal cancer, and gastric cancer.25,26
PD-L1
Next, we have PDL-1. Immune cells need a way to differentiate between normal cells and cells that are dangerous to the body. This is accomplished through “checkpoint” proteins such as PD-L1. When PD-L1 on a tumor cell combines with PD-1 on an immune cell, it prevents the immune cells from killing the tumor cell.27 Tumor cells sometimes take advantage of checkpoints such as these to avoid being killed and proliferate.28,29
T Cells
T cells play a critical role in immune response to not only pathogenic infections, but cancer as well.30 There are two major types of T cells: CD8+ cells and CD4+ cells:21
Epithelial to Mesenchymal Transition (EMT)
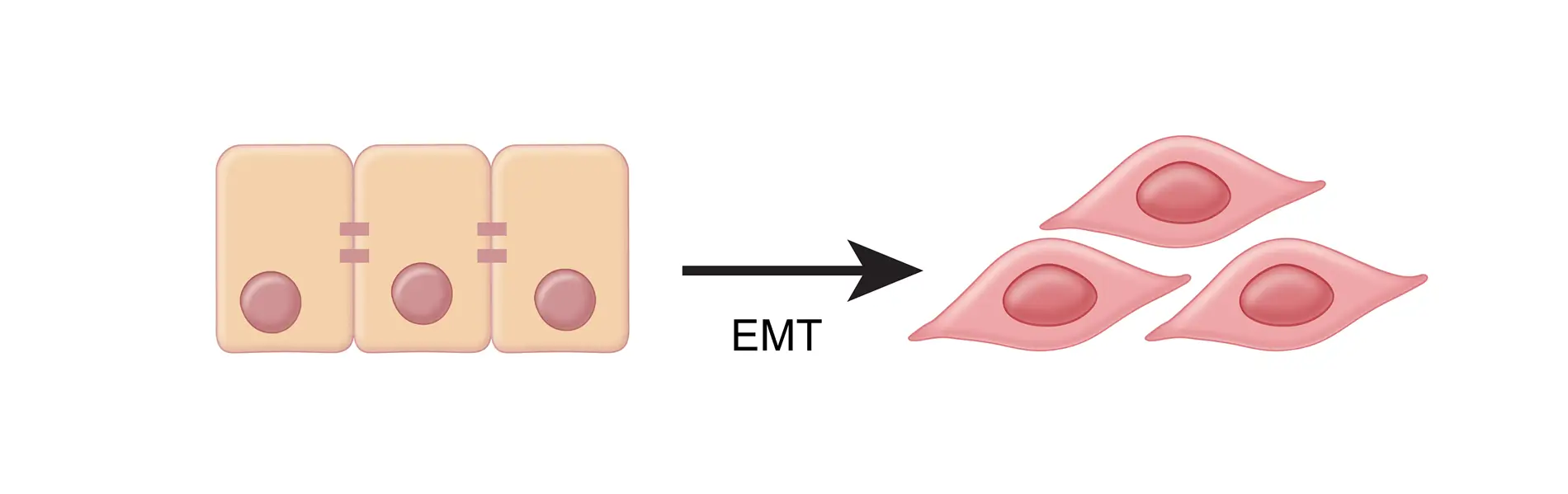
Next, we have the Epithelial to Mesencymal Transition process, or “EMT”. Similar to how TGF-β converts fibroblasts into harmful CAF’s, EMT converts epithelial cells to more motile mesenchymal cells.9 EMT is an important property of cancer cells that allows them to become invasive, metastatic, and migratory.38,39
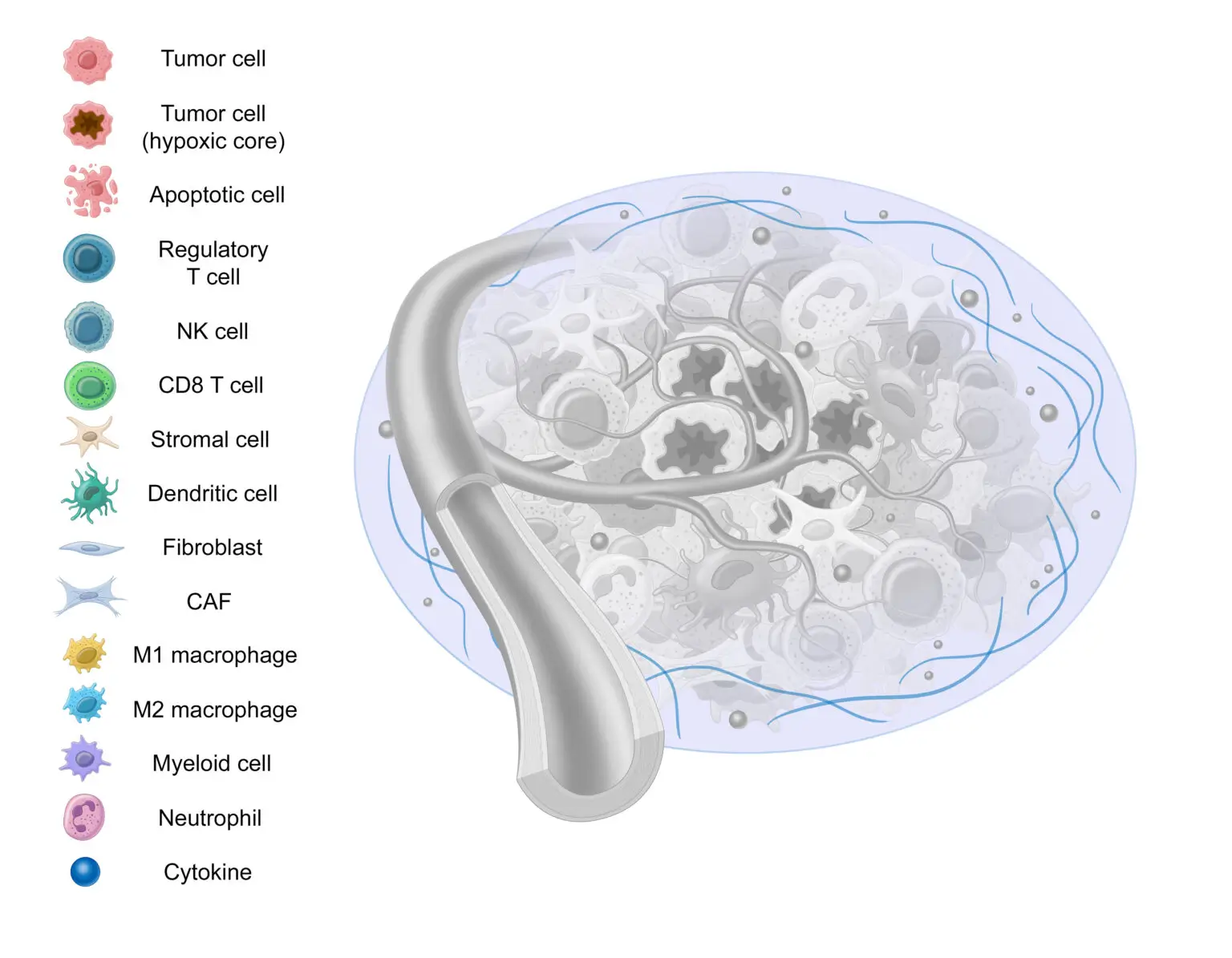
Lastly, we have the Extracellular Matrix (ECM), a highly dynamic structure present in all tissues that’s comprised of collagen, elastin, fibronectin, and other structural proteins.11,40 Serving as a form of scaffolding, the ECM serves as a physical barrier, provides structure, and helps cells attach and communicate with other cells.41,42
The ECM undergoes constant, controlled remodeling.40 Unfortunately, this same remodeling can contribute to tumor progression and resistance to therapy.43
Conclusion
Cancer development is incredibly complex, and often is not caused by a single mutation. This is why Afirma GSC uses whole-transcriptome derived analysis: it better allows us to capture the full complexity of cancer and generate a wealth of valuable information. This information not only serves as the foundation for Afirma GSC, but also creates the potential for novel research. If you’re interested in conducting research of your own or would like to learn more about Afirma GSC, fill out the form below and a genomic specialist will be in touch with you shortly.
You may have seen references to “BRAF-like”, “RAS-like”, or “Non-BRAF-Non-RAS ” alterations in the Afirma GSC patient report. Shown below is the Results Summary section of a patient report with NRAS p.Q61R detected. Note how it describes the tumor as exhibiting a “RAS-like profile”:

But what does this mean? BRAF-like, RAS-like, and Non-BRAF-Non-RAS are three different molecular profiles that describe different types of tumor behavior.
Cancer Genome Atlas Network, Cell, 2014.
The BRAF-like and RAS-like profiles were first conceived in 2014, when the Cancer Genome Atlas developed a 71-gene signature to classify alterations into one of two categories: as having either “BRAF– like” or “RAS-like” gene expression.1 This was done by investigating 391 Papillary Thyroid Cancers, which often have two mutually exclusive cancer drivers (BRAFV600E and mutated RAS) and assigning a BRAFV600E – RAS Score (BRS).
Yoo et al, PLoS Genetics, 2016.
Yoo et al expanded on this 2 years later to include follicular thyroid cancers, follicular adenomas, and add one more molecular subtype: “non-BRAF-non-RAS” (NBNR).2
Tang AL, et al, Endocrine Practice, 2020.
The table below shows how several genomic alterations would be organized into the different molecular classes:3
Tumor behavior by molecular profile
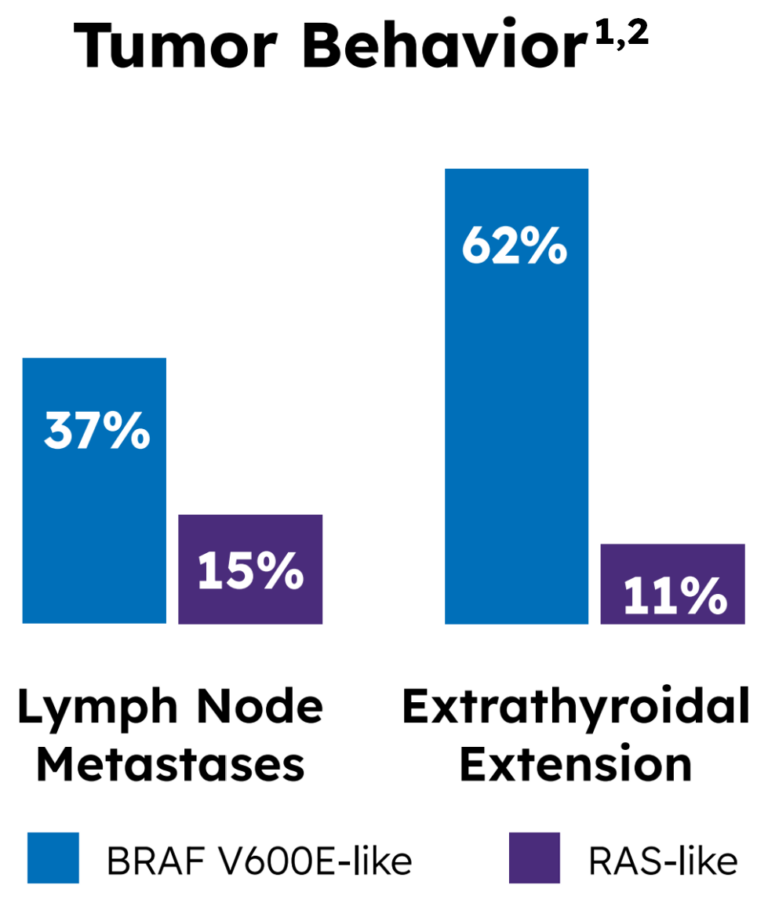
Yoo also provided insight into potential tumor behavior: a higher frequency of lymph node metastases was found in 37% of the BRAF-like group, compared to 15% of the RAS-like group. Similarly, a higher frequency of extrathyroidal extension was found in 62% of the BRAF-like group, compared to 11% of the RAS-like group.1,2
It’s very important to note, however, that none of the values above reach 100%. While the different molecular profiles tend to correlate to different tumor behaviors, this is at a population level that does not take individual variability into account.
Conclusion
Cancer development is incredibly complex, and often is not caused by a single mutation. This is one of the reasons why Afirma GSC uses whole-transcriptome-derived analysis: it allows us to go beyond the variants to characterize nodules further. If you’re interested in conducting research of your own or would like to learn more about Afirma GSC, fill out the form below and a genomic specialist will be in touch with you shortly.
To run the Afirma Genomic Sequencing Classifier (or GSC), Veracyte uses a dedicated FNA sample. This is because a fresh FNA sample is the proven standard for molecular testing.1 Though some may view slide scraping as a convenient way to obtain genomic material for molecular testing, there are significant limitations to this collection methodology.
Lupo et al. Diagnostic Cytopathology 2020.
First, obtaining an adequate amount of genomic material can be a challenge.
In the validation study of a molecular test touting the use of slide scraping authored by Lupo et al, 18% of samples failed to provide adequate genomic material.2
Nikiforova et al. Cancer Cytopathology 2020.
A rate of 19% was seen in another molecular test’s validation study authored by Nikiforova et al.3 Of greater concern, however, is the discordance of detection of molecular alterations between slide scraping and obtaining a fresh sample via FNA. In the Nikiforova publication, 17% of gene fusions and 14% of copy number alterations were missed on slide scraping relative to a matched fresh sample.3
In fact, the discussion section of this manuscript states:
The collection of a portion of a fresh FNA sample directly into a nucleic acid preservative solution should be attempted whenever possible because this provides the highest success rate and accuracy of testing.”
Cancer Cytopathology 2020
Conclusion
Given these limitations, Afirma GSC is only available to order for use on a dedicated FNA pass.
To learn more about implementing Afirma GSC or obtain a new test requisition form, get in touch with a Veracyte Genomic Specialist using the form below.
In the field of cancer diagnostics, precision is paramount. This is where Veracyte’s Afirma Genomic Sequencing Classifier® (GSC) steps in, leveraging the power of whole-transcriptome derived analysis to provide actionable insights into thyroid nodule tumor management. But what exactly is the transcriptome, and how does the Afirma GSC leverage it to help providers and their patients?
Simply put, the word “transcriptome” is the combination of the words “transcript” and “genome”. To fully understand what the transcriptome is and the role that it plays, we need to start at the DNA and end at the protein. This flow of genetic information from DNA to RNA, and then finally to the protein, is referred to as “the central dogma.”
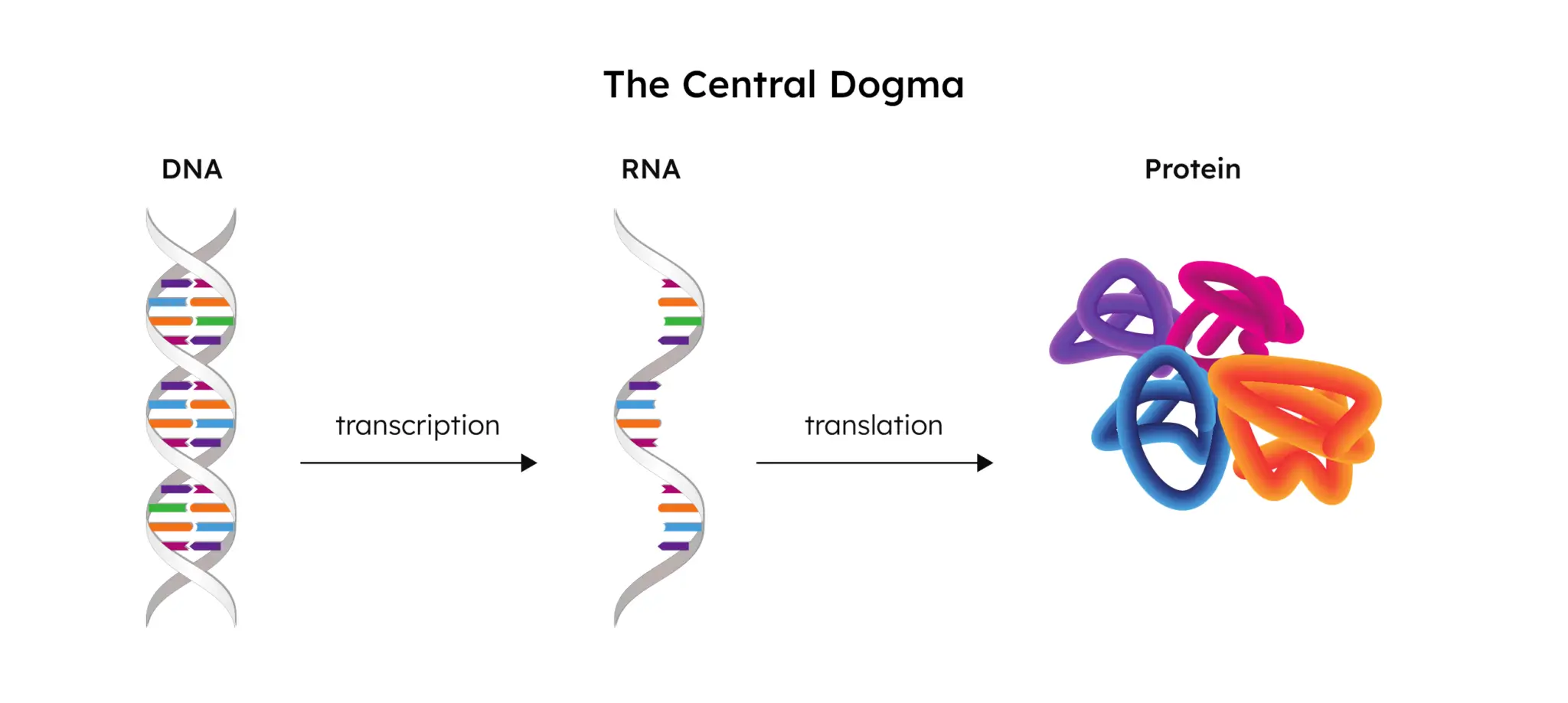
Transcription
The human genome is made up of DNA, the blueprint used to create all of the cells in an organism. When a gene from this DNA is activated (“expressed”), a portion of the DNA is unzipped by RNA polymerase and an RNA copy is created.
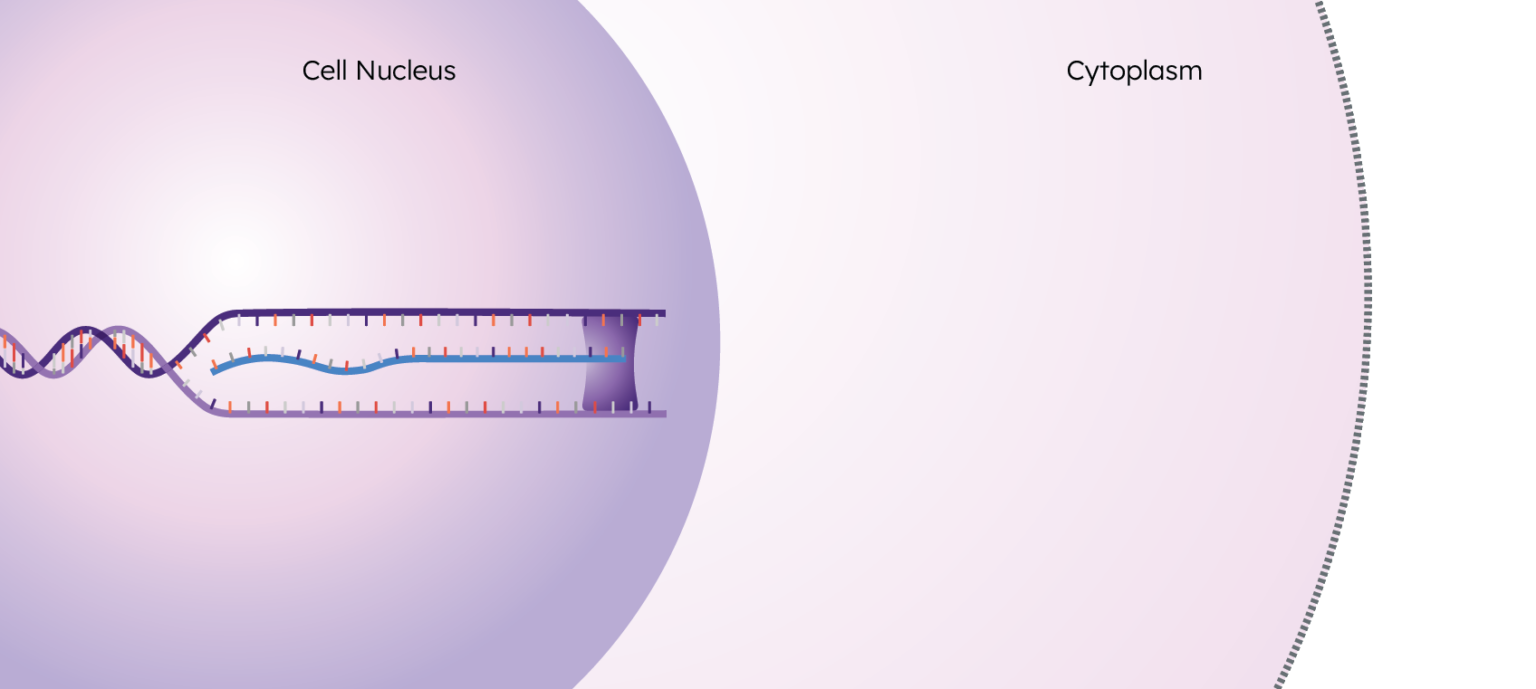
These RNA copies are called “transcripts”, and this process of creating them is called “transcription”. These transcripts exit the nucleus and enter the cytoplasm to eventually undergo translation, resulting in the final step of the central dogma: the protein.
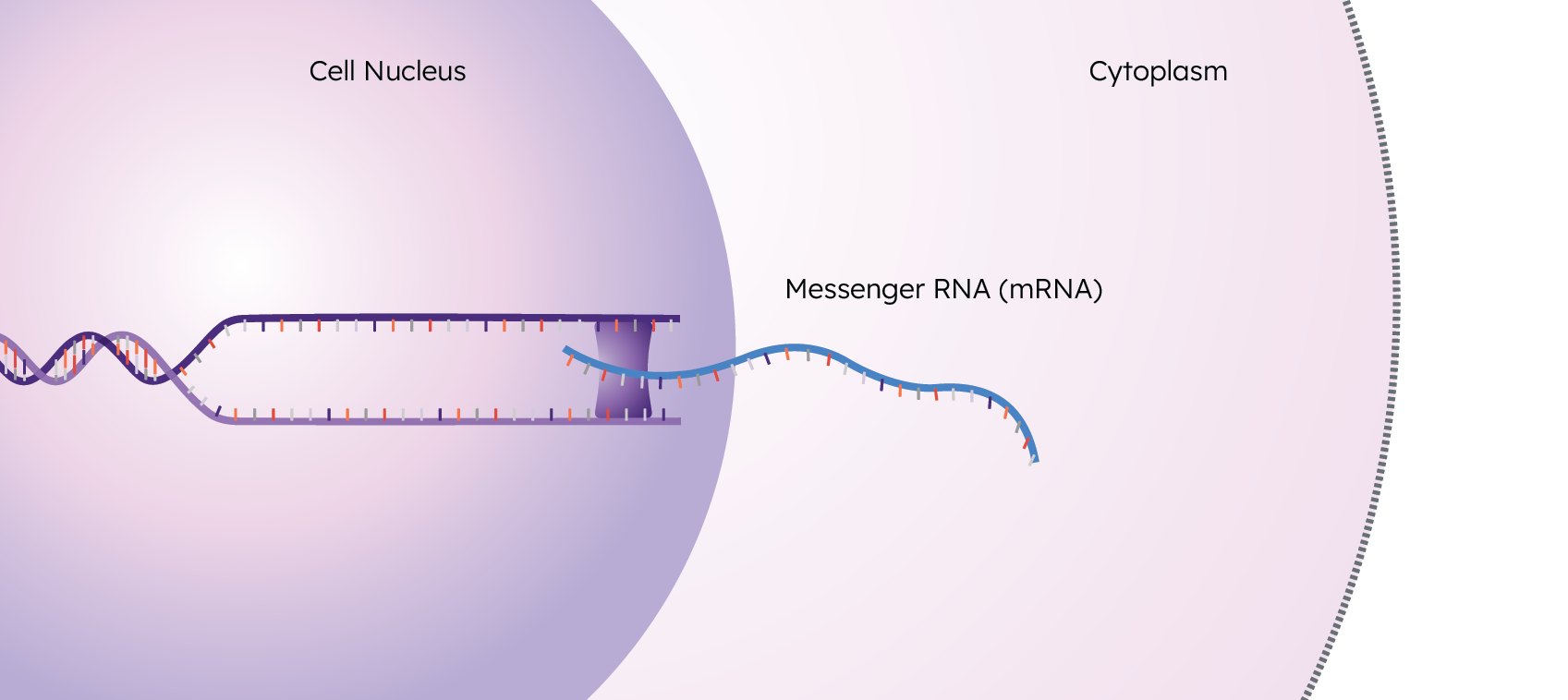
The collection of all of these transcripts in a sample is the transcriptome.
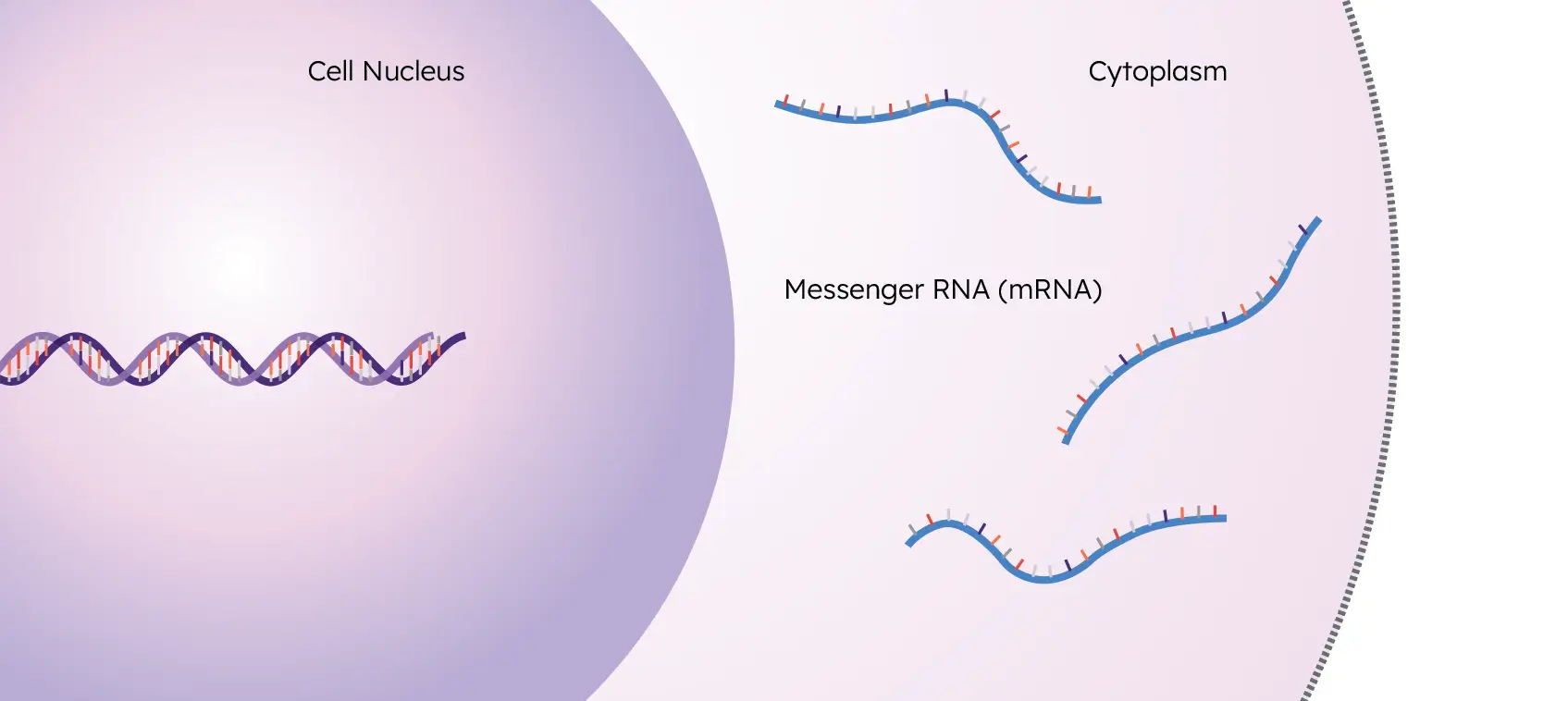
What the Transcriptome Tells Us
As the functional output of the genome, the transcriptome can provide valuable information that the genome can’t. As mentioned earlier, in order for a transcript to exist, a gene must get activated – this is called “gene expression”. By looking at which transcripts are present in a cell, we can see which genes are actually getting expressed – and by looking at how many transcripts are present, we get a quantitative measure of how expressed they are.
This is important, because the presence of a genetic variant does not guarantee its expression. Heart and brain cells feature the same exact DNA, for instance, and yet are entirely different in appearance and function due to mRNA expression. Also, the presence of an alteration may or may not be relevant: some mutations are cancer drivers, some are low-risk, and some are not yet understood.
Assessing the genome provides valuable insight into what alterations are present in a tumor, but in order to understand how the tumor behaves (or how the genome behaves), you have to assess the transcriptome. This cannot be accomplished by looking at DNA alone.
Mitochondrial Transcripts & Loss of Heterozygosity
The genome also misses critical information provided by mitochondrial transcripts due to the nuclear genome being separate from the mitochondrial genome. With whole-transcriptome derived analysis, Afirma GSC is able to leverage both mitochondrial transcripts and chromosomal loss of heterozygosity to classify oncocytic (Hürthle) cell lesions with high sensitivity.1,2
DNA analysis provides valuable information, but it’s not always clinically actionable for these reasons. Since the transcriptome is further along the central dogma and closer to the end product (the protein), it can provide a better picture of what’s actually going on within a cell.
Conclusion
Cancer development is incredibly complex, and often is not caused by a single mutation. This is why Afirma GSC uses whole-transcriptome derived analysis: it better allows us to capture the full complexity of cancer and generate a wealth of valuable information. This information not only serves as the foundation for Afirma GSC, but also creates the potential for novel research. If you’re interested in conducting research of your own or would like to learn more about Afirma GSC, fill out the form below and a genomic specialist will be in touch with you shortly.
Oncocytic cells, also known as Hürthle cells, are present in more than 20% of indeterminate thyroid nodules and are often challenging to classify as high or low risk.1,2,3 Due to an abundance of oncocytic cells, both oncocytic carcinomas and adenomas share a similar cytologic appearance.4 Therefore, diagnosis has historically relied on identifying capsular and vascular invasion, which can be challenging to diagnose in instances of minimally invasive disease.5
To address this challenge, Veracyte’s Afirma Genomic Sequencing Classifier (GSC) includes two algorithmic components that are able to classify oncocytic cell lesions with high sensitivity.1,6
First is the Oncocytic Cell Index, which uses mRNA expression and mitochondrial transcripts to identify fine needle aspiration (FNA) samples with a genomic signature associated with oncocytic features. Oncocytes contain more mitochondria than normal follicular cells, and mitochondria have their own genome that is separate from the nuclear genome.5 The RNA sequencing performed by Veracyte allows us to capture and quantify from a molecular perspective if patient samples are oncocytic or not.
The second index is the Oncocytic Neoplasm Index, which leverages mRNA expression and chromosomal loss of heterozygosity (LOH) to identify if a nodule with oncocytic cells is also molecularly neoplastic. Research has shown that oncocytic neoplasms have higher levels of LOH than other subtypes.7
The result of these two indices is an 89% sensitivity in classifying oncocytic cell lesions which is important to know because about three out of four oncocytic cell lesions are classified as benign.1,2,3,6,7,8,9,10 Together, these indices have helped improve the specificity and PPV of the benign vs. suspicious call that the Afirma GSC ensemble classifier provides.1,2,9,10 The performance of Afirma GSC in this subset of challenging nodules has not only been demonstrated in Afirma GSC’s validation study, but has been confirmed in multiple other studies as well.1,2,6,7,9,10 By addressing the challenge in diagnosing this particular neoplasm, Veracyte aims to improve outcomes for patients.