Out of the Xpression Atlas database of 177,227 thyroid nodule FNA samples, 382 samples were identified with BRAF fusions in this study (99% of which were reported as Afirma GSC Suspicious). These 382 samples were analyzed by Bethesda category, fusion partner genes, and 5 molecular signatures of aggressiveness.
Out of the entire database of both BRAF fusion positive and negative nodules, the BRAF fusion positive nodules constituted:
- 0.1% of Bethesda III nodules
- 0.33% of Bethesda IV nodules
- 1.21% of Bethesda V nodules
- 1.18% of Bethesda VI nodules
This study identified 75 different partner genes, the most frequent of which were:
- SND1 (25%)
- AGK (19%)
- MKRN1 (10%)
- WARS (3%)
- EXOC4 (2%)
- TRIM24 (2%)
- ZC3HAV1 (2%)
The 5 genomic signatures of aggressiveness used to assess the samples included:
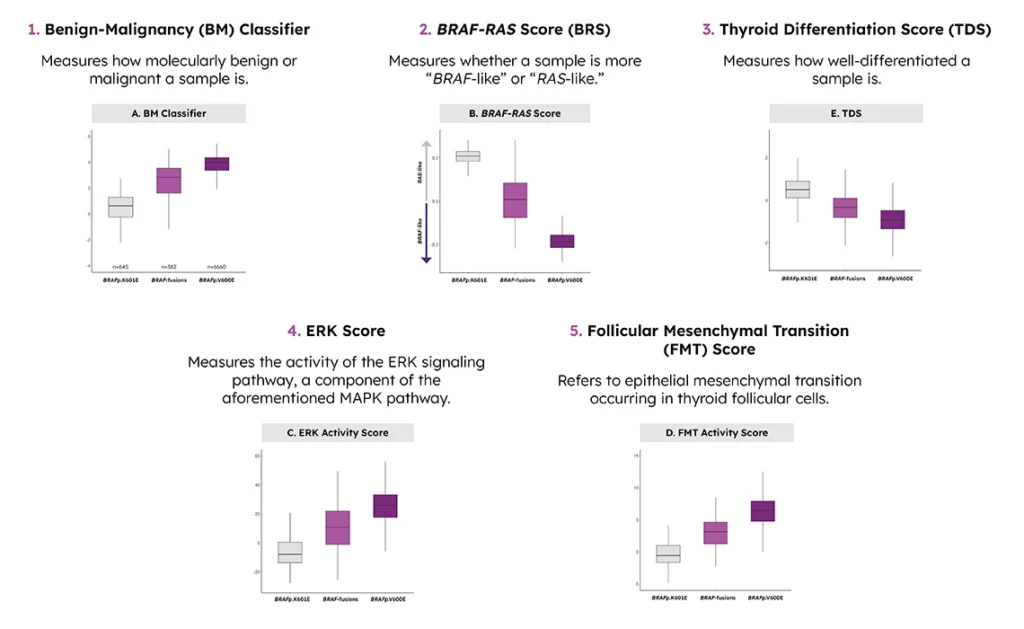
Using these signatures, the BRAF fusion samples were compared to samples with BRAFp. K601E alterations (RAS-like) and samples with BRAFp.V600E alterations (BRAF-like). This analysis showed that with all 5 signatures, BRAF fusions had expression levels between tumors with more aggressive BRAFp.V600E alterations and tumors with less aggressive BRAFp.K601E alterations.
Conclusion
We’re happy that Veracyte could contribute to the meaningful research presented at the Association for Molecular Pathology (AMP) 2024 Annual Meeting & Expo. If you’d like to learn about how Afirma is leveraging whole-transcriptome-derived analysis to create the potential for novel research opportunities, contact us here, and someone from our team will be in touch with you shortly.
Conference Materials
Afirma Thyroid
Assessment of BRAF Fusions in 177,227 Thyroid Nodules by Exome-Enriched RNA-Seq Testing
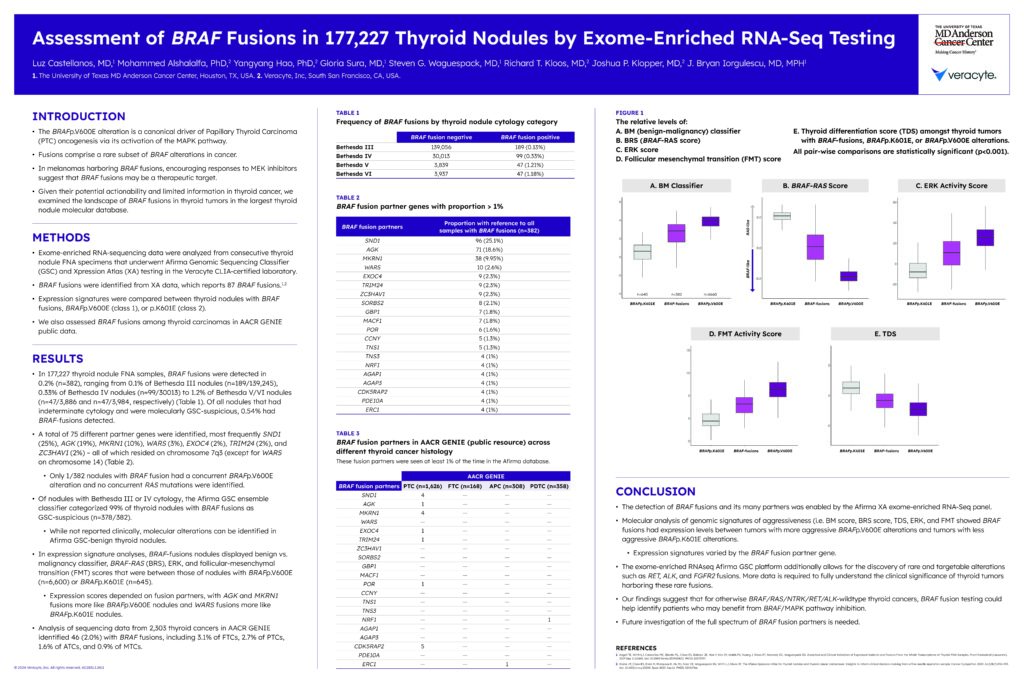

Castellanos L, et al. AMP. 2024.